DENGUE Virus
1. Genome Organization & Viral Protein Expression
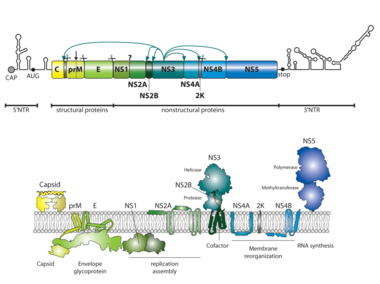
DV belongs to the genus Flavivirus in the family Flaviviridae. These viruses have a single stranded RNA genome of positive polarity, which is about 11,000 nucleotides long. The RNA carries a single open reading frame encoding a polyprotein that is translated in a cap-dependent manner at the rough endoplasmic reticulum (rER). The viral polyprotein is co- and post-translationally processed into three structural proteins (C, prM, and E) and seven non-structural (NS) proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5). The amino termini of prM, E, NS1 and NS4B are generated upon cleavage by the host ER signal peptidase in the lumen of the ER, whereas the processing of most of the other NS proteins and the C-terminus of the C protein is carried out by the viral two-component protease NS2B-NS3 in the cytoplasm of DV infected cells. For the cleavage of the C-terminus of NS1 an unknown ER peptidase seems to be responsible, and a Golgi localized furin protease cleaves prM at a later state of infection to generate the mature form of M.
DV RNA replication occurs in close association with cellular membranes that may serve as a scaffold for the viral replication complexes (RC). In fact, virus infection induces invaginations of the ER membrane that are connected to the cytoplasm via around 11nm-diameter pores allowing the influx of metabolites required for RNA replication and the release of progeny RNA genomes. NS5 was identified as the RNA-dependent RNA polymerase (RdRp) and has also a methyltransferase activity important for the formation of the RNA cap-structure and for 2’-O-methylation. NS3 acts as the viral serine protease, which requires the protein NS2B as cofactor for full enzymatic activity and to tether the protein to intracellular membranes. In addition, NS3 also comprises an RNA triphosphatase as well as an RNA helicase with nucleoside triphosphatase activity. NS1 is also required for viral RNA replication by contributing to the formation of the membranous replication complexes. In addition, NS1 is secreted and contributes to viral pathogenesis. NS2A was reported to allow the transfer of viral RNA from the replicase to assembling virions, while NS4A and NS4B play a critical role in RC formation. In addition, both proteins might serve to anchor the viral RC to membranes. Moreover, it is assumed that these proteins contribute to the inhibition of the host interferon response.
The genome organization is shown on the top. 5’ and 3’ non-translated regions (NTRs) are indicated with stem-loop structures. Cleavage sites of the viral NS2B-3 protease are indicated with arrows. Scissors refer to cleavage sites targeted by the host cell signal peptidase. Membrane topologies of viral cleavage products are given on the bottom, along with functions ascribed to the proteins.
< Overview Page 2 (Replication Cycle) >