Die AG Guizetti im Detail
Aktuelle Mitarbeiter*innen
Dr. Julien Guizetti (AG-Leiter, seit 2017)
Sultan Bekbayev (MSc-Arbeit, seit 2023)
Anja Klemmer (Doktorandin, seit 2019)
& wechselnde BSc Studierende
Alumni
Yannik Voß (MSc Arbeit, 2018; Doktorand, 2019 - 2023)
Nicolas Lichti (BSc Arbeit, 2019; MSc Arbeit, 2022 - 2023)
Christoph Wenz (MSc Praktikum, 2021; MSc Arbeit, 2022 - 2023)
Vanessa Stürmer (Praktikantin, 2018-2019; MSc Praktikum, 2020 - 2021; MSc Arbeit, 2022 - 2023)
Sophie Stopper (MSc Praktikum, 2022)
Caroline Simon (Doktorandin, 2018 - 2022)
Marius Flörchinger (BSc Arbeit, 2021; MSc Praktikum, 2021)
Johanna Bauer (BSc Arbeit, 2020; MSc Praktikum, 2021)
Dr. Tatiany Patricia Romão (Gastwissenschaftlerin, 2020 - 2021)
Nathan Ribot (MSc Praktikum, 2020)
Ann-Kathrin Mehnert (MSc Praktikum, 2018; MSc Arbeit, 2019)
Marlen Wesemeyer (BSc Arbeit, 2018)
Nadja Ilkenhans (BSc Arbeit, 2018)
Forschungsschwerpunkt
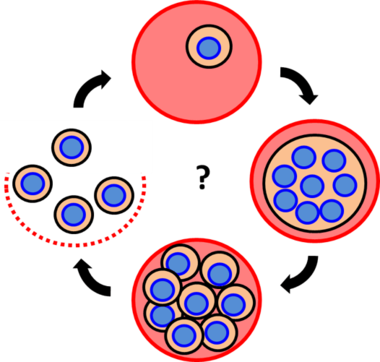
Der Schweregrad der Malaria-Erkrankung, an der immer noch etwa 600.000 Menschen pro Jahr sterben, wird durch die Anzahl der mit Plasmodium infizierten roten Blutkörperchen, die im Blutkreislauf zirkulieren, bestimmt. Dennoch haben wir nur ein rudimentäres Verständnis über den unkonventionellen Zellteilungsmechanismus, den der Parasit für seine rasche Vermehrung nutzt. Im Gegensatz zu den meisten Modellorganismen, die sich durch binäre Spaltung teilen, vervielfältigen Malaria verursachende Parasiten zunächst ihre Zellkerne und erzeugen später 8-30 Tochterzellen auf einmal (Abb. 1). I
In der AG Guizetti untersuchen wir den Mechanismus der Kernteilung, um letztlich herauszufinden, wie der Parasit seine endgültige Anzahl von Tochterzellen erreicht. Wir setzen moderne Technologien der Superauflösungs-, Elektronen- und Lebendzellmikroskopie in Kombination mit CRISPR/Cas9-Genom-Editierung ein, um die Kinetik der Kernvermehrung zu beschreiben, die Organisation und Zusammensetzung des Zentrosoms zu untersuchen und die mutmaßliche Rolle der Phasentrennung bei der Funktion des Zentrosoms aufzudecken (siehe Projekte). Durch die Untersuchung dieses abweichenden Eukaryoten tragen wir zu einer evolutionären zellbiologischen Perspektive bei, wollen aber auch neue Ziele innerhalb dieses essentiellen Signalwegs aufdecken, die im Kampf gegen Malaria genutzt werden können.
-

In "einfachen" Worten:
Malaria ist eine schwere Krankheit, an der immer noch mehr als ein halbe Million Menschen jährlich versterben. Die Krankheit entsteht während sich der Malaria-Erreger, genannt Plasmodium falciparum, im Blut eines infizierten Menschen massiv vermehrt. Nach einem infektiösen Mückenstich und einem Zwischenstopp in der Leber dringt der kleine Erreger in unsere roten Blutzellen ein. Dort verdaut er die Blutzelle von innen und wächst. In der zweiten Hälfte dieses Entwicklungszyklus beginnt er seine eigenen Zellkerne, die das Erbmaterial in Form von DNA enthalten, zu vermehren. Erst zum Schluss seiner Entwicklung verpackt er die Zellkerne in 8 bis 30 Tochterzellen, die dann aus der platzenden Wirtszelle entkommen, um dann viele neue rote Blutzellen zu infizieren. Dieser Vermehrungsprozess unterscheidet sich signifikant von dem, der sich in z.B. unseren Körperzellen abspielt, die üblicherweise eine Zweiteilung durchlaufen. Auch deshalb verstehen wir die Zellteilung des Parasiten recht wenig. Ziel der AG Guizetti ist es mithilfe von modernen Mikroskopietechniken diesen Prozess mit einer hohen zeitlichen und räumlichen Auflösung zu untersuchen und zu verstehen (siehe Filme und Bilder). Wir hoffen, damit neue Zielmoleküle für Interventionsstrategien im Kampf gegen Malaria aufzudecken.
Mehr Infos zu Malaria gibt es in unserem Service Bereich!
Projekte
Die Zellteilung ist ein hochdynamischer Prozess. Im Falle von Malariaparasiten wurden die wichtigsten zellulären Parameter jedoch noch nicht zeitaufgelöst untersucht. Wir entwickeln Zeitraffer-Bildgebungsverfahren in Kombination mit Färbeprotokollen für lebende Zellen, um den Zeitpunkt der Teilung, die Kernvermehrungsraten und die Reorganisation der Spindel und mehr auf der Ebene einzelner Parasiten zu bestimmen (Abb. 2). Auf diese Weise schaffen wir einen quantitativen zellbiologischen Rahmen, der die Untersuchung der Zellteilung von Malariaparasiten mit einer noch nie dagewesenen räumlichen und zeitlichen Auflösung ermöglichen wird.
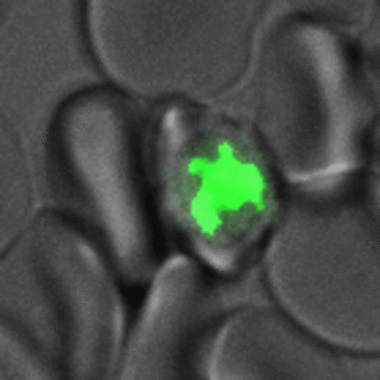
Abbildung 2. Malaria-Parasiten produzieren mehrere Kerne, bevor sie die Wirtszelle verlassen. Zeitrafferaufnahme einer von einem Parasiten infizierten roten Blutzelle vom Beginn der Kernteilung bis zum Aufplatzen und Wiedereintritt in die Nachbarzelle. Die rote Blutzelle und der Parasit sind im Übertragungskanal (grau) zu sehen, während das Chromatin, d. h. die Kerne, des Parasiten durch die Expression von GFP-markiertem Histon 2B (grün) fluoresziert sind.
Die Kernteilung erfordert kleine und komplizierte molekulare Maschinen, die die Chromosomen nach ihrer Replikation physisch auseinanderziehen. Schlüsselkomponenten dieser Maschinerie sind i) die mitotische Spindel, die aus röhrenförmigen Polymeren, den Mikrotubuli, besteht, an denen einzelne Chromosomen befestigt sind, und ii) die Zentrosomen, die die Mikrotubuli organisieren und die beiden Pole bilden, zu denen die Chromosomen gezogen werden (Abb. 3). Die Organisation der überraschend kleinen mitotischen Spindel und des atypischen Zentrosoms in Plasmodium ist noch nicht gut untersucht worden. Mit einer Kombination aus Superresolution und Elektronenmikroskopie haben wir das erste detaillierte Arbeitsmodell des Zentrosoms des Malariaparasiten erstellt (Abb. 3). Anhand dieses Modells können wir nun Schritt für Schritt die Komponenten dieser divergenten Organelle identifizieren und ihre Funktion aufklären.
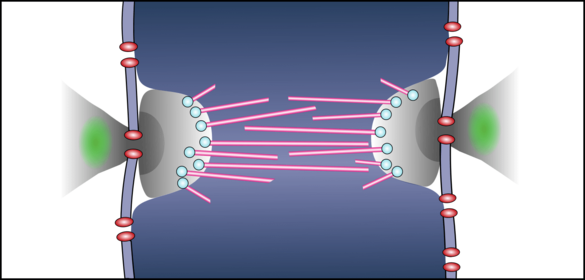
Abbildung 3. Arbeitsmodell des Zentrosoms und der mitotischen Spindel des Malariaparasiten. Mikrotubuli (magenta) bilden eine bipolare Anordnung, die mitotische Spindel. Der Spindelpol, auch Zentrosom (grau) genannt, ist eine proteinreiche Struktur, die die Kernmembran (graublau) durch eine spezielle Kernpore (rot) überspannt. Der extranukleäre Teil enthält Zentrine (grün), eine der wenigen konservierten zentrosomalen Proteinfamilien. Der intranukleare Teil beherbergt die Mikrotubuli-Nukleationsstelle (weiß), von der aus sich die Mikrotubuli in Richtung der Kern-DNA (blau) ausbreiten. Später wird sich diese mitotische Spindel ausdehnen, um die Schwesterchromosomen auseinanderzutreiben. Angepasst aus Simon et al. 2021 (PMID: 34535568).
Zentrosomen können als membranlose Organellen betrachtet werden, die viele verschiedene Komponenten in einem bestimmten subzellulären Kompartiment konzentrieren. In den letzten Jahren wurde das neue biophysikalische Konzept der Phasentrennung, das die Neigung bestimmter Proteine zur Koazervierung unter bestimmten Bedingungen beschreibt, zur Erklärung der Eigenschaften von membranlosen Organellen, einschließlich der Zentrosomen, herangezogen. Die Phasentrennung wurde bei Malaria verursachenden Parasiten noch nicht untersucht, aber bei der In-vitro-Untersuchung rekombinanter P. falciparum-Zentrine, bei denen es sich um konservierte kalziumbindende Proteine handelt, stellten wir fest, dass einige von ihnen eine Flüssig-Flüssig-Phasentrennung erfahren können (Abb. 4). Die Bedeutung dieser Eigenschaft für den Aufbau und die Funktion des Zentrosoms in vivo ist Gegenstand weiterer Untersuchungen der Gruppe.
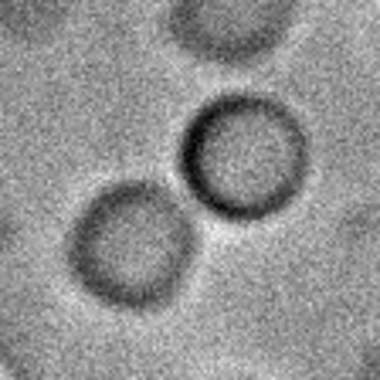
Abbildung 4. Rekombinante Centrin-Protein-Tröpfchen zeigen ein flüssigkeitsähnliches Verhalten. Transmissions-Zeitraffer-Mikroskopie einer rekombinanten PfCen1-Proteinlösung nach Zugabe von Kalzium. Die Fusion zwischen zwei Proteintröpfchen deutet darauf hin, dass es sich eher um Flüssig-Flüssig-Phasenkondensate als um feste Proteinansammlungen handelt. Angepasst von Voss et al. 2022 (biorxiv.org/content/10.1101/2022.07.26.501452v1).
Film-Galerie
Film 1 - Malariaparasiten erzeugen mehrere Tochterzellen, die in benachbarte Erythrozyten eindringen.
Film 2 - Hochaufgelöste dreidimensionale Organisation von Mikrotubuli und Zentrosomen in mehrkernigen Malariaparasiten.
Weitere Filme
Film 3 - Mehrere Runden mitotische Spindelbildung und Ausdehnung Malariaparasit.
Film 5 - Dreidimensionale Organisation von Mikrotubuli, Kernporen und mitotischer Spindel in sich teilenden Kernen mit Hilfe der Ultrastrukturexpansionsmikroskopie.
Film 4 - Dreidimensionale Organisation von Mikrotubuli, zentriolären Plaques, Kernporen und Hemispindeln in sich teilenden Kernen mittels Ultrastrukturexpansionsmikroskopie.
Publikationen / Kollaborationen
Originalarbeiten
Voß Y, Klaus S, Lichti NP, Ganter M, Guizetti J (2023) Malaria parasite centrins can assemble by Ca2+-inducible condensation. PLoS Pathog. Dec 27;19(12):e1011899. doi: 10.1371/journal.ppat.1011899. PMID: 38150475; PMCID: PMC10775985.
Machado M, Klaus S, Klaschka D, Guizetti J, Ganter M (2023) Plasmodium falciparum CRK4 links early mitotic events to the onset of S-phase during schizogony. mBio. Jun 22:e0077923. doi: 10.1128/mbio.00779-23. Epub ahead of print. PMID: 37345936.
Wenz C, Simon CS, Romão TP, Stürmer VS, Machado M, Klages N, Klemmer A, Voß Y, Ganter M, Brochet M, Guizetti J (2023) An Sfi1-like centrin-interacting centriolar plaque protein affects nuclear microtubule homeostasis. PLoS Pathog. May 2;19(5):e1011325. doi: 10.1371/journal.ppat.1011325. PMID: 37130129; PMCID: PMC10180636.
Ganter M*, Guizetti J*, Kilian N* (2022) Visualization of Infected Red Blood Cell Surface Antigens by Fluorescence Microscopy In: Jensen ATR, Hviid L, editors. Malaria Immunology: Targeting the Surface of Infected Erythrocytes. New York, NY: Springer US. pp. 425–433. doi:10.1007/978-1-0716-2189-9_31
Voss Y, Klaus S, Ganter M, Guizetti J* (2022) Ca2+-inducible phase separation of centrins in proliferating malaria parasites. bioRxiv 2022.07.26.501452. doi:10.1101/2022.07.26.501452
Klaus S, Binder P, Kim J, Machado M, Funaya C, Schaaf V, Klaschka D, Kudulyte A, Cyrklaff M, Laketa V, Höfer T, Guizetti J, Becker NB, Frischknecht F, Schwarz US, Ganter M (2022) Asynchronous nuclear cycles in multinucleated Plasmodium falciparum facilitate rapid proliferation. Sci Adv. Apr;8(13):eabj5362. doi: 10.1126/sciadv.abj5362. Epub 2022 Mar 30. PMID: 35353560.
Machado M, Klaus S, Klaschka D, Guizetti J, Ganter M (2022) Plasmodium falciparum CRK4 links early mitotic events to the onset of S-phase during schizogony. bioRxiv 2022.08.31.505163. doi:10.1101/2022.08.31.505163.
Wenz C, Simon CS, Romão TP, Stürmer V, Machado M, Klages N, Klemmer A, Voß Y, Ganter M, Brochet M, Guizetti J* (2022) An Sfi1-like centrin-interacting centriolar plaque protein affects nuclear microtubule homeostasis. bioRxiv 2022.07.28.501831. doi:10.1101/2022.07.28.501831
Diehl M, Roling L, Rohland L, Weber S, Cyrklaff M, Sanchez CP, Beretta CA, Simon CS, Guizetti J, Hahn J, Schulz N, Mayer MP, Przyborski JM (2021) Co-chaperone involvement in knob biogenesis implicates host-derived chaperones in malaria virulence. PLoS Pathog. Oct 6;17(10):e1009969. doi: 10.1371/journal.ppat.1009969. PMID: 34614006; PMCID: PMC8544838.
Simon CS, Funaya C, Bauer J, Voβ Y, Machado M, Penning A, Klaschka D, Cyrklaff M, Kim J, Ganter M, Guizetti J (2021) An extended DNA-free intranuclear compartment organizes centrosome microtubules in malaria parasites. Life Sci Alliance. Sep 17;4(11):e202101199. doi: 10.26508/lsa.202101199. PMID: 34535568; PMCID: PMC8473725.
Simon CS, Stürmer VS, Guizetti J (2021) How Many Is Enough? - Challenges of Multinucleated Cell Division in Malaria Parasites. Front Cell Infect Microbiol. May 7;11:658616. doi: 10.3389/fcimb.2021.658616. PMID: 34026661; PMCID: PMC8137892.
Soni K, Kempf G, Manalastas-Cantos K, Hendricks A, Flemming D, Guizetti J, Simon B, Frischknecht F, Svergun DI, Wild K, Sinning I (2021) Structural analysis of the SRP Alu domain from Plasmodium falciparum reveals a non-canonical open conformation. Commun Biol. May 20;4(1):600. doi: 10.1038/s42003-021-02132-y. PMID: 34017052; PMCID: PMC8137916.
Guizetti J, Frischknecht F (2021) Apicomplexans: A conoid ring unites them all. PLoS Biol. Mar 11;19(3):e3001105. doi: 10.1371/journal.pbio.3001105. PMID: 33705378; PMCID: PMC7951970.
Barcons-Simon A, Cordon-Obras C, Guizetti J, Bryant JM, Scherf A. (2020) CRISPR Interference of a Clonally Variant GC-Rich Noncoding RNA Family Leads to General Repression of var Genes in Plasmodium falciparum. mBio. Jan 21;11(1)
Mehnert, A.K., Simon C.S., Guizetti, J. (2019). Immunofluorescence staining protocol for STED nanoscopy of Plasmodium-infected red blood cells. Mol Biochem Parasitol. 229, 47–52.
Müller, L. S. M., Cosentino, R. O., Förstner, K. U., Guizetti, J., Wedel, C., Kaplan, N., … Siegel, T. N. (2018). Genome organization and DNA accessibility control antigenic variation in trypanosomes. Nature 563(7729), 121–125
Zanghì, G., Vembar, S. S., Baumgarten, S., Ding, S., Guizetti, J., Bryant, J. M., … Scherf, A. (2018). A Specific PfEMP1 Is Expressed in P. falciparum Sporozoites and Plays a Role in Hepatocyte Infection. Cell Reports 22(11), 2809–2817.
Bryant J.M., Regnault C., Scheidig-Benatar C., Baumgarten S., Guizetti J.*, Scherf A. (2017). CRISPR/Cas9 Genome Editing Reveals That the Intron Is Not Essential for var2csa Gene Activation or Silencing in Plasmodium falciparum. MBio 8(4). pii: e00729-17.
Guizetti, J.*, Barcons-Simon, A., Scherf, A. (2016). Trans-acting GC-rich non-coding RNA at var expression site modulates gene counting in malaria parasite. Nucleic Acids Res 44, 9710–9718.
Zhang, Q., Siegel, T. N., Martins, R. M., Wang, F., Cao, J., Gao, Q., Cheng, X., Jiang, L., Hon, C. C., Scheidig-Benatar, C., Sakamoto, H., Turner, L., Jensen, A. T., Claes, A., Guizetti, J., Malmquist, N. A., and Scherf, A. (2014). Exonuclease-mediated degradation of nascent RNA silences genes linked to severe malaria. Nature 513, 431–435.
Guizetti, J., Martins, R. M., Guadagnini, S., Claes, A., and Scherf, A. (2013). Nuclear Pores and Perinuclear Expression Sites of var and Ribosomal DNA Genes Correspond to Physically Distinct Regions in Plasmodium falciparum. Eukaryot Cell 12, 697–702.
Guizetti J*, Scherf A. (2013). Silence, activate, poise and switch! Mechanisms of antigenic variation in Plasmodium falciparum. Cell Microbiol. doi:10.1111/cmi.12115.
Guizetti J, Gerlich DW. (2012). ESCRT-III polymers in membrane neck constriction. Trends Cell Biol 22:133–140. doi:10.1016/j.tcb.2011.11.007.
Guizetti, J., Schermelleh, L., Mantler, J., Maar, S., Poser, I., Leonhardt, H., Muller-Reichert, T., and Gerlich, D. W. (2011). Cortical constriction during abscission involves helices of ESCRT-III-dependent filaments. Science 331, 1616–1620.
Guizetti, J., Mantler, J., Muller-Reichert, T., and Gerlich, D. W. (2010). Correlative time-lapse imaging and electron microscopy to study abscission in HeLa cells. Methods Cell Biol 96, 591–601.
Lacroix, B., van Dijk, J., Gold, N. D., Guizetti, J., Aldrian-Herrada, G., Rogowski, K., Gerlich, D. W., and Janke, C. (2010). Tubulin polyglutamylation stimulates spastin-mediated microtubule severing. J Cell Biol 189, 945–954.
Guizetti J, Gerlich DW. (2010). Cytokinetic abscission in animal cells. Semin Cell Dev Biol 21:909–916. doi:10.1016/j.semcdb.2010.08.001.
Guizetti J, Mäntler J, Müller-Reichert T, Gerlich DW. (2010). Correlative time-lapse imaging and electron microscopy to study abscission in hela cells. Methods Cell Biol 96:591–601. doi:10.1016/S0091-679X(10)96024-X.
Lekomtsev, S., Guizetti, J., Pozniakovsky, A., Gerlich, D. W., and Petronczki, M. (2010). Evidence that the tumor-suppressor protein BRCA2 does not regulate cytokinesis in human cells. J Cell Sci 123, 1395–1400.
Steigemann, P., Wurzenberger, C., Schmitz, M. H., Held, M., Guizetti, J., Maar, S., and Gerlich, D. W. (2009). Aurora B‑mediated abscission checkpoint protects against tetraploidization. Cell 136, 473–484.
Reviews
Voß Y, Klaus S, Guizetti J, Ganter M. (2023) Plasmodium schizogony, a chronology of the parasite's cell cycle in the blood stage. PLoS Pathog. Mar 2;19(3):e1011157. doi: 10.1371/journal.ppat.1011157. PMID: 36862652; PMCID: PMC9980825.
Guizetti, J.*, and Scherf, A. (2013). Silence, activate, poise and switch! Mechanisms of antigenic variation in Plasmodium falciparum. Cell Microbiol 15, 718–726.
Guizetti, J., and Gerlich, D. W. (2012). ESCRT-III polymers in membrane neck constriction. Trends Cell Biol 22, 133–140.
Guizetti, J., and Gerlich, D. W. (2010). Cytokinetic abscission in animal cells. Semin Cell Dev Biol 21, 909–916.
Externe Kollaborationspartner
Mathieu Brochet, University of Geneva
Paul Guichard and Virginie Hamel, University of Geneva
Robert Moon, London School of Hygiene and Tropical Medicine
Moritz Treeck, Francis Crick Institute London
Kai Johnsson, MPI for Medical Research Heidelberg
Sebastian Baumgarten and Jessica Bryant, Institut Pasteur Paris
Group activities
News - AG Guizetti
Artikel in "Life Science Alliance"
(Veröffentlichung: 17. September 2021)
In diesem Artikel stellen das Guizetti-Labor und Kollegen ein erstes Modell des atypischen Parasiten-Zentrosoms vor, welches für die Steuerung der schnellen Zellteilung des Malaria-Erregers Plasmodium unerlässlich ist:
"Augenblick für Forschung"
(Online-Vortrag unseres AG-Leiters Julien)
Im Rahmen der monatlichen Seminarreihe "Augenblick für Forschung" der Daimler und Benz Stiftung berichtet unser AG-Leiter Julien in einerm 15-minütigen Online-Vortrag mit dem Titel "Malaria-Erreger im Blut – Vermehren und Verstecken" zu seiner Forschung. Die Reihe bietet Einblicke in aktuelle Forschungsthemen, die von der Daimler und Benz Stiftung gefördert werden. Die Protagonisten sind Stipendiaten des Postdoktoranden-Förderprogramms, die ihre individuellen Projekte vorstellen. Es gibt also jeden Monat etwas Spannendes zu entdecken!