Helmholtz-University Junior Research Group “Cell Plasticity and Epigenetic Remodeling”
Head: Dr. Darjus Tschaharganeh
Facilitated by advances in genome sequencing technologies during the past 30 years researchers are now able to obtain a comprehensive overview of genomic alterations in almost every cancer type. These data revealed a number of frequent alterations and mutational patterns in different cancer types, pinpointing to potential candidate “driver” genes for particular diseases. Although this information is very valuable it is purely descriptive. Therefore, subsequent functional studies in appropriate model systems are necessary to assign phenotypic consequences to these “drivers”, which not only help to understand cancer biology but also facilitate development of new therapeutic targets for personalized cancer medicine. Our laboratory has strong expertise in applying functional genomic tools, such as RNA-interference and CRISPR/Cas9, to create innovative and powerful mouse models of cancer. We use these tools to generate genetically defined tumors, which closely resemble alterations found in human cancer samples, thereby probing the function of specific genetic events on tumor development. Subsequently we utilize the models we create for investigating specific vulnerabilities created by these defined genetic events by employing state-of-the-art high-throughput screening technology in vitro and in vivo.
Our future research will continue to characterize cancer genomes using the functional genomics approaches outlined above. In particular we are interested in epigenetic modifications as well as genes responsible for mediating epigenetic remodeling. This group of genes is frequently altered in human cancer but remains poorly characterized. We aim to investigate their relevance in cancer development and to define their suitability to represent specific vulnerabilities for cancer therapy. In fact a variety of epigenetic drugs are currently under development and show promising success. Furthermore, our lab will develop technologies to interrogate epigenomes in vitro and in vivo to facilitate the study of epigenetic processes. More recently we are interested to adjust our mouse model technology for studying the impact of the immune system in tumorigenesis. Here our research will focus on constructing synthetic genetic circuits for visualizing and interrogating intercellular communication of tumor cells and immune cells as well as defining determinants for improving immunotherapeutic responses
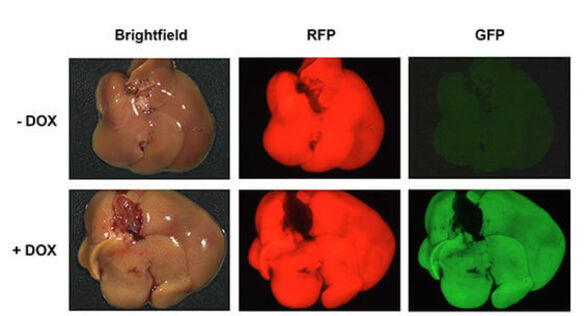
Liver-specific shRNA transgenic mice as a tool to interrogate gene function in vivo. Dissection-microscope pictures of livers from shRNA transgenic animals harboring a Doxycycline (DOX) inducible shRNA construct for potent target gene suppression. shRNA expression is coupled to a GFP-reporter. Mice receiving Dox-chow (+ DOX) strongly express GFP and the shRNA construct as well as RFP as a second reporter allele specifically in the liver. Livers of mice on normal diet (- DOX) only express the RFP reporter.
Team
Group Leader: Dr. Darjus Tschaharganeh
Email: d.tschaharganeh(at)dkfz.de
Administration: Brigitte Brauch
Postdocs:
- Dr. Marco Breinig
- Dr. Katrin Ganzenberg
- Dr. Stephan Krieg
- Dr. Hendrik Wiethoff
Students:
- Peter Macsek
- Agnieszka Seretny
- Sonia Jiménez-Vázquez
- Anne Landsmann
Technicians:
- Luise Butthof
- Lio Böse
- Lena Wendler-Link
Funding

Publications
2017
Kastenhuber ER, Lalazar G, Houlihan SL, Tschaharganeh DF, Baslan T, Chen CC, Requena D, Tian S, Bosbach B, Wilkinson JE, Simon SM, Lowe SW. DNAJB1-PRKACA fusion kinase interacts with β-catenin and the liver regenerative response to drive fibrolamellar hepatocellular carcinoma. Proc Natl Acad Sci U S A. 2017 Nov 21.
Wan S, Meyer AS, Weiler SME, Rupp C, Tóth M, Sticht C, Singer S, Thomann S, Roessler S, Schorpp-Kistner M, Schmitt J, Gretz N, Angel P, Tschaharganeh DF, Marquardt J, Schirmacher P, Pinna F, Breuhahn K. Cytoplasmic localization of the cell polarity factor Scribble supports liver tumor formation and tumor cell invasiveness. Hepatology. 2017 Nov 20.
Wall MA, Shaffer TM, Harmsen S, Tschaharganeh DF, Huang CH, Lowe SW, Drain CM, Kircher MF. Chelator-Free Radiolabeling of SERRS Nanoparticles for Whole-Body PET and Intraoperative Raman Imaging. Theranostics. 2017 Jul 22;7(12):3068-3077.
Méndez-Lucas A, Li X, Hu J, Che L, Song X, Jia J, Wang J, Xie C, Driscoll PC, Tschaharganeh DF, Calvisi DF, Yuneva M, Chen X. Glucose Catabolism in Liver Tumors Induced by c-MYC Can Be Sustained by Various PKM1/PKM2 Ratios and Pyruvate Kinase Activities. Cancer Res. 2017 Aug 15;77(16):4355-4364.
Yuan D, Huang S, Berger E, Liu L, Gross N, Heinzmann F, Ringelhan M, Connor TO, Stadler M, Meister M, Weber J, Öllinger R, Simonavicius N, Reisinger F, Hartmann D, Meyer R, Reich M, Seehawer M, Leone V, Höchst B, Wohlleber D, Jörs S, Prinz M, Spalding D, Protzer U, Luedde T, Terracciano L, Matter M, Longerich T, Knolle P, Ried T, Keitel V, Geisler F, Unger K, Cinnamon E, Pikarsky E, Hüser N, Davis RJ, Tschaharganeh DF, Rad R, Weber A, Zender L, Haller D, Heikenwalder M. Kupffer Cell-Derived Tnf Triggers Cholangiocellular Tumorigenesis through JNK due to Chronic Mitochondrial Dysfunction and ROS. Cancer Cell. 2017 Jun 12;31(6):771-789.e6.
Pelossof R, Fairchild L, Huang CH, Widmer C, Sreedharan VT, Sinha N, Lai DY, Guan Y, Premsrirut PK, Tschaharganeh DF, Hoffmann T, Thapar V, Xiang Q, Garippa RJ, Rätsch G, Zuber J, Lowe SW, Leslie CS, Fellmann C. Prediction of potent shRNAs with a sequential classification algorithm. Nat Biotechnol. 2017 Apr;35(4):350-353.
2016
Tasdemir N, Banito A, Roe JS, Alonso-Curbelo D, Camiolo M, Tschaharganeh DF, Huang CH, Aksoy O, Bolden JE, Chen CC, Fennell M, Thapar V, Chicas A, Vakoc CR, Lowe SW. BRD4 Connects Enhancer Remodeling to Senescence Immune Surveillance. Cancer Discov. 2016 Jun;6(6):612-29.
Tschaharganeh DF#, Bosbach B#, Lowe SW. (# equal contribution) Coordinated Tumor Suppression by Chromosome 8p. Cancer Cell. 2016 May 9;29(5):617-9.
Tschaharganeh DF, Lowe SW, Garippa RJ, Livshits G. Using CRISPR/Cas to study gene function and model disease in vivo. FEBS J. 2016 May 5.
2015
Dow LE, O'Rourke KP, Simon J, Tschaharganeh DF, van Es JH, Clevers H, Lowe SW. APC restoration promotes cellular differentiation and reestablishes crypt homeostasis in colorectal cancer. Cell. 2015 Jun 18;161(7):1539-52.
Dow LE, Fisher J, O'Rourke KP, Muley A, Kastenhuber ER, Livshits G, Tschaharganeh DF, Socci ND, Lowe SW. Inducible in vivo genome editing with CRISPR-Cas9. Nat Biotechnol. 2015 Apr;33(4):390-4.
2014
Huang CH, Lujambio A, Zuber J, Tschaharganeh DF, Doran MG, Evans MJ, Kitzing T, Zhu N, de Stanchina E, Sawyers CL, Armstrong SA, Lewis JS, Sherr CJ, Lowe SW. CDK9 mediated transcriptional elongation is required for Myc addiction in hepatocellular carcinoma. Genes Dev. 2014 Aug 15,28(16):1800-14
Tschaharganeh DF, Xue W, Calvisi DF, Evert M, Michurina TV, Dow LE, Banito A, Katz SF, Kastenhuber ER, Weissmueller S, Huang CH, Lechel A, Andersen JB, Capper D, Zender L, Longerich T, Enikolopov G, Lowe SW p53 dependent Nestin regulation links tumor suppression to cellular plasticity in liver cancer. Cell. 2014 Jul 31;158(3):579-92.
Weissmueller S, Manchado E, Saborowski M, Morris JP 4th, Wagenblast E, Davis CA, Moon SH, Pfister NT, Tschaharganeh DF, Kitzing T, Aust D, Markert EK, Wu J, Grimmond SM, Pilarsky C, Prives C, Biankin AV, Lowe SW. Mutant p53 drives pancreatic cancer metastasis through cell-autonomous PDGF receptor β signaling. Cell. 2014 Apr 10;157(2):382-94.
2013
Tschaharganeh DF, Chen X, Latzko P, Malz M, Gaida MM, Felix K, Ladu S, Singer S, Pinna F, Gretz N, Sticht C, Tomasi ML, Delogu S, Evert M, Fan B, Ribback S, Jiang L, Brozzetti S, Bergmann F, Dombrowski F, Schirmacher P, Calvisi DF, Breuhahn K. Yes-associated protein up-regulates Jagged-1 and activates the Notch pathway in human hepatocellular carcinoma. Gastroenterology. 2013 Jun;144(7):1530-1542.e12.
Lujambio A, Akkari L, Simon J, Grace D, Tschaharganeh DF, Bolden JE, Zhao Z, Thapar V, Joyce JA, Krizhanovsky V, Lowe SW. Non-cell-autonomous tumor suppression by p53. Cell. 2013 Apr 11;153(2):449-60.