Research Projects
Establishment and Exploitation of a European-Latin American Research Consortium towards Eradication of Preventable Gallbladder Cancer - EULAT Eradicate GBC
Funded by the European Union's Horizon 2020 research and innovation programme under grant agreement No 825741
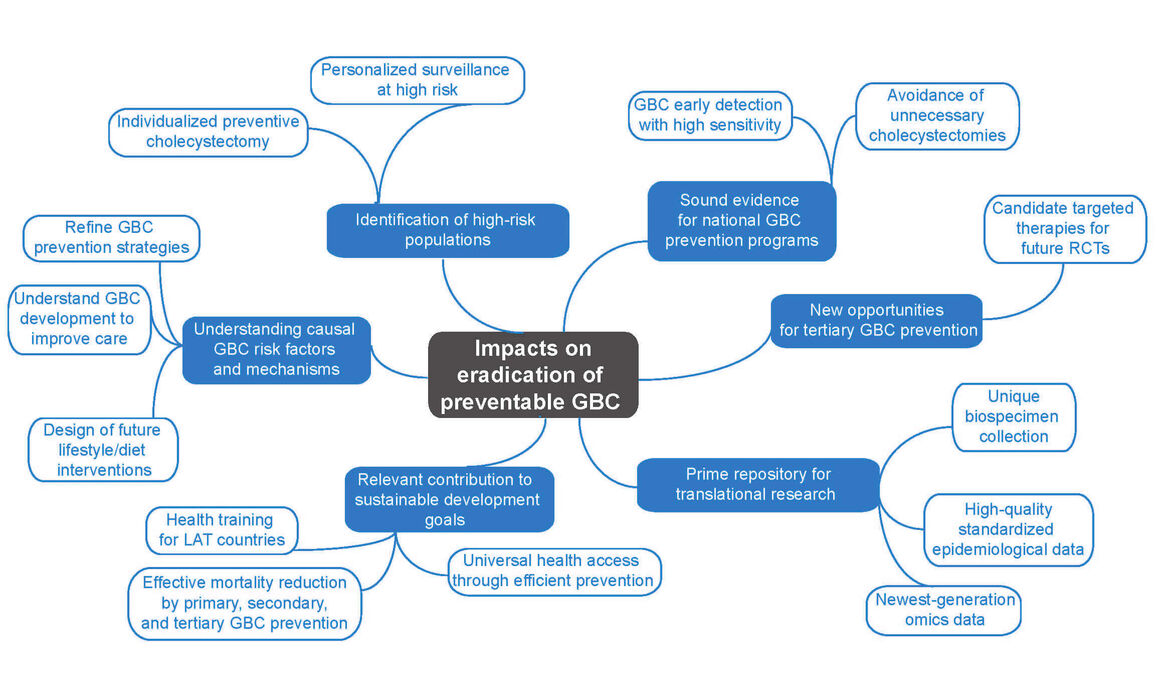
Gallbladder cancer (GBC) is a neglected disease with huge potential for prevention. This project aims at significantly improving the accuracy of risk estimation and early detection of GBC by identifying and adequately considering geographical, environmental, lifestyle, ethnic, gender and molecular differences. We plan to generate the information needed to establish and refine current prevention programmes, including the primary, secondary and tertiary prevention of GBC. We will (1) build a unique European–Latin American GBC biorepository integrated into a tailored IT platform, (2) identify, validate and functionally characterize novel GBC biomarkers, (3) develop a multifactorial risk score that integrates established and newly identified epidemiological and molecular risk factors, (4) improve the understanding of the causal mechanisms that link lifestyle, cultural and behavioural factors to GBC development, (5) unravel novel opportunities for the targeted therapy of incidental GBC, (6) exploit existing and newly generated epidemiological and multi-omics data to improve the accuracy of GBC risk prediction and (7) contribute to the training of the next generation of Latin American researchers in precision medicine for GBC. The generated information will permit identification of individuals at high GBC risk, guiding surveillance and individual decisions on the possible benefit of preventive gallbladder removal in regions of low and high GBC incidence. Novel data on genomic alterations in incidental GBC will pave the way towards implementation of future clinical trials. The planned European–Latin American GBC biorepository and IT platform will constitute a prime resource for translational research on individualized prevention, personalized early detection and targeted therapy of GBC. The participation in our project of representatives of health authorities, patients and the industry guarantees the efficient incorporation of project results into national health policies.
Please see also:
https://cordis.europa.eu/project/id/825741
Publications:
https://pubmed.ncbi.nlm.nih.gov/33020926/
Identification and validation of circulating sncRNAs causally associated with gallbladder cancer and development of a multifactorial risk prediction score
Funded by the German Research Foundation (DFG). For a video with preliminary results please click here
Summary
Our project aims at investigating the causal mechanisms that underlie gallbladder cancer development, and at developing and validating a risk score for gallbladder cancer prediction before clinical manifestation, eventually revolutionizing the prevention of this neglected disease.
The objectives of the project are:
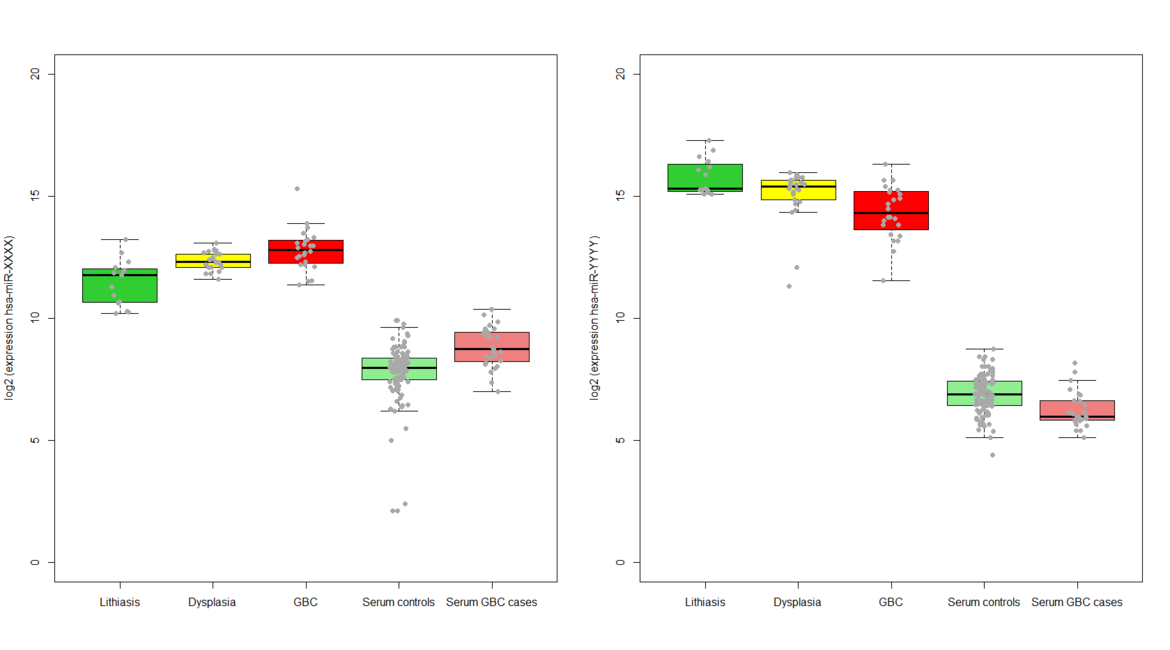
1) To pre-select candidate small non-coding RNAs based on expression changes in gallbladder tissue along the multistage model of carcinogenesis represented by the sequence gallstones and inflammation ® dysplasia ® gallbladder cancer
2) To identify and validate circulating small non-coding RNAs associated with gallbladder cancer risk
3) To test the causality and to estimate the causal effects of associated small non-coding RNAs by Mendelian randomization
4) To establish a risk score which combines established gallbladder cancer risk factors and newly identified small non-coding RNAs associated with gallbladder cancer risk
Towards Personalized Treatment of Chagas Disease by Molecular Profiling of Patients and Parasites EULACH16/T020108
Funded by ERANet LAC
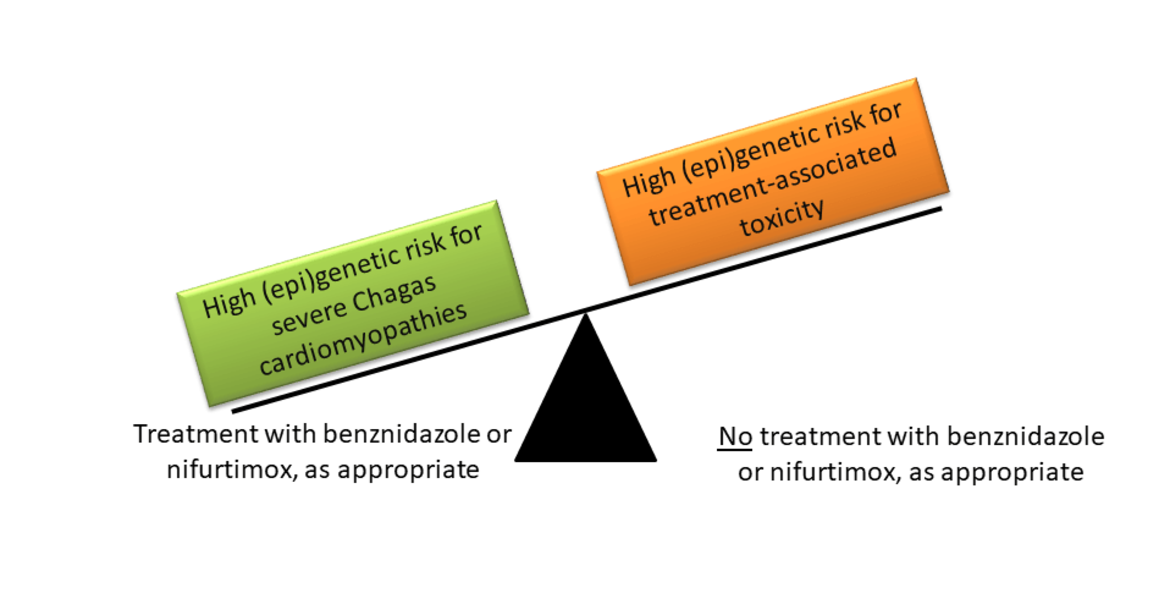
Chagas disease is a potentially fatal illness caused by the parasite Trypanosoma cruzi. There are more than 6 million people infected worldwide, mainly in endemic areas of Latin America, but also increasingly in Europe due to migration and travel. Recent achievements in Chagas prevention include a better control of vector, and a substantial reduction of infections via blood transfusion and mother to child transmission. However, enhanced treatment of millions of infected people remains a key challenge. Infected persons can be treated with benznidazole (BZ) or nifurtimox (NFX). BZ and NFX show close to 100% effectivity in killing the parasite shortly after infection, but efficacy sharply decreases with post infection time. This decline in effectivity represents an important limitation because symptoms are often absent or unspecific during the initial, acute phase of Chagas disease. Treatment may be indicated during the chronic phase to prevent disease progression reducing risk of cardiac, digestive and neurological complications, but treatment is also accompanied by adverse reactions, which occur in up to 40% of treated patients. A personalized treatment of Chagas disease based on the (epi)genetic makeup of patients and parasites would permit to optimize the benefit-risk ratio of chronic infected patients, simultaneously reducing treatment costs.
Using a three stage approach, this project will quantify the impact of (epi)genetic variability on (1) BZ and NFX treatment efficiency and treatment-related toxicities, (2) the prognosis of Chagas disease (e.g. cardiomyopathy development), and (3) Chagas susceptibility and associated deaths. Human genetic and epigenetic variability (characterized by genomewide genotypes and miRNA profiles in pretreatment serum), the genetic makeup of T. cruzi (characterized by DTUs), and possible interactions among them will be investigated.
Our innovative research project will deliver new insights to use molecular information (1) to predict whether Chagas treatment in the chronic phase will be beneficial or rather detrimental, (2) to predict the course of Chagas disease and guide patient management, and (3) to identify geographical regions and minority ethnic groups particularly threatened by Chagas disease. This information will help to improve the management of Chagas disease not only in Latin America but in the whole world.
Mendelian Randomization
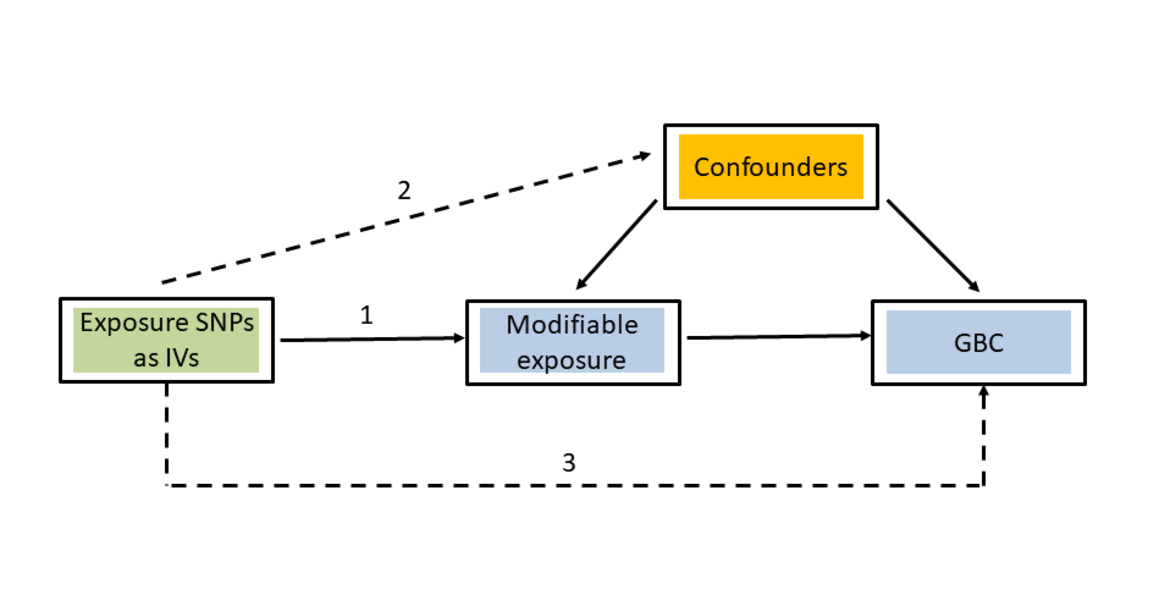
Mendelian randomization (MR) is an analytical method used to assess the causal effects of particular modifiable risk factors (exposures) on specific phenotypes (outcomes). The rationale for MR studies lies in the use of genetic variants as instrumental variables (IV). Genetic variants are randomly assigned at conception, mimicking the design of a randomized controlled trial. The use of genetic variants strongly associated with the investigated modifiable risk factor then guarantees that causal effect estimates from MR analyses are less likely to be affected by confounding or reverse causation than the results of traditional observational studies. The selection of instrumental variables is essential to perform a valid MR study. In order to be able to test the causal effect of a modifiable risk factor (e.g. BMI in the present study) on an outcome (e.g. GBC in the present study), the selected genetic variants must fulfil the following three criteria:
(1) Strong association with the modifiable risk factor
(2) No association with any confounder
(3) Association with the outcome exclusively through the investigated modifiable risk factor
Fulfilment of the first criterion can be guaranteed by selecting instrumental variables associated with the investigated modifiable risk factor at a genome-wide level of statistical significance. MR studies that do not fulfil this criterion often translate into low statistical power and potentially biased results. Since checking the second and third criteria is more difficult, alternative methods that are relatively robust against violations of these conditions have been developed, such as MR-Egger regression, weighted median and mode-based MR estimates.
Publications:
Robust Statistics in Molecular Medicine
The results of genetic association studies are often influenced by observations deviating from the majority of the data - so-called outliers. We are examining the benefits and limitations of robust regression models and develop new robust statistical methods to robustify the results of genetic association studies. In particular, we applied the bounded Huber and extended the R package “robustbase” with the re-descending Hampel functions to down-weight outlier influence. Furthermore, we conducted comprehensive simulations and analyzed real protein, metabolite, mRNA expression and genotype data to compare the prediction accuracy of standard and robust Huber-LASSO.
Publications:
Collaborative research projects
Novel Breast Cancer Susceptibility Variants
In collaboration with the Research Group “Molecular Genetics of Breast Cancer” at the German Cancer Research Center (Head Prof. Ute Hamann, http://www.dkfz.de/en/mammakarzinom/index.php), we have investigated the relationship between genetic variability in the chromosomal passenger complex and the risk of breast cancer. Fifteen polymorphisms in INCENP, AURKB, BIRC5 and CDCA8 were genotyped as part of the Collaborative Oncological Gene-environment Study (COGS). Possible associations were investigated in a fixed-effects model meta-analysis comprising 88,911 European women from 39 case-control studies of the Breast Cancer Association Consortium (BCAC).
Publications:
IMPACT
In collaboration with the Division of Preventive Oncology at the German Cancer Research Center (Head Prof. Hermann Brenner, https://www.dkfz.de/en/praeventive-onkologie/index.php), we investigate the association of genetic variants and gene expression profiles with colorectal cancer risk and disease outcome.
Publications: