Epigenomics, Epitranscriptomics and novel therapy approaches in AML

Our vision - what we do
The aim of our group is to identify pathogenetic mechanisms in acute leukemias and other cancers to ultimately develop and test novel therapy approaches in early phase clinical trials. Clinical specimens from these trials are then analyzed to further understand mechanisms of disease and therapy resistance. Group leaders are Maximilian Blank (MD), Maike Janssen (MD), Carsten Müller-Tidow (MD, Professor of Medicine), Moritz Cornelius Pauli (MD) and Fengbiao Zhou. Senior scientists of our group are Stefanie Göllner, Yi Liu and Christian Rohde, lead investigator Marina Scheller. For the clinical trials we work closely with the different groups in the Department and the NCT.
Who We Are
Group Leaders
Senior Scientist
Postdocs
PhD Students MMPU (EMBL and Heidelberg University Hospital)
Biologielaboranten/-innen
Technische Assistentin
Wiss. Koordinatorin
Scientific Interest
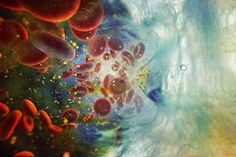
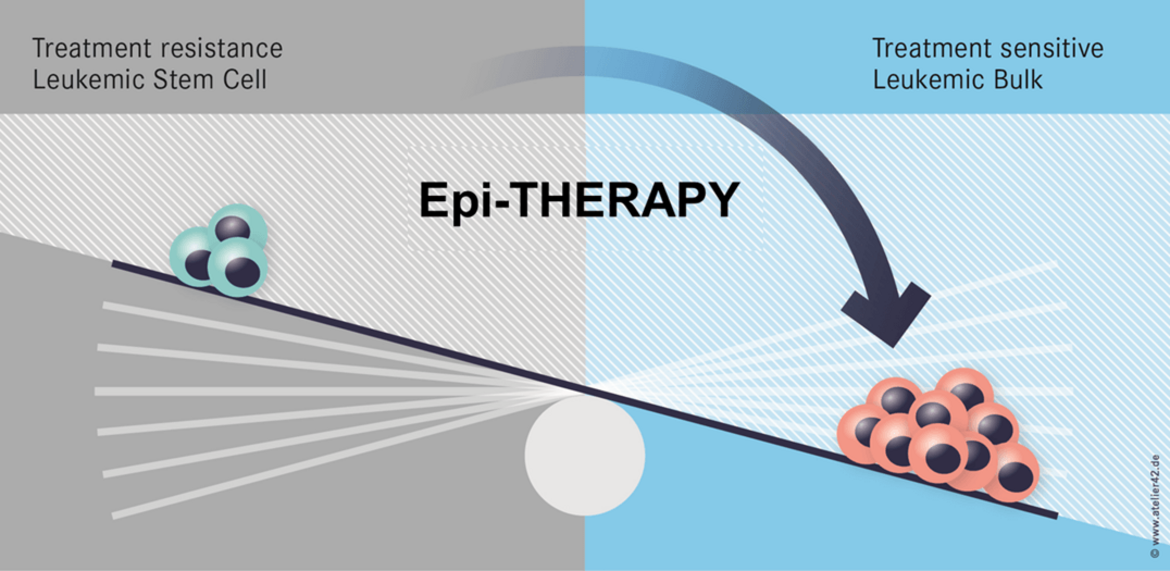
One important hypothesis of our work states that epigenetic changes affect stemness of cancer cells and cause therapy resistance. Hence, epigenetic therapies may be an efficient approach to overcome therapy resistance (Figure 1).
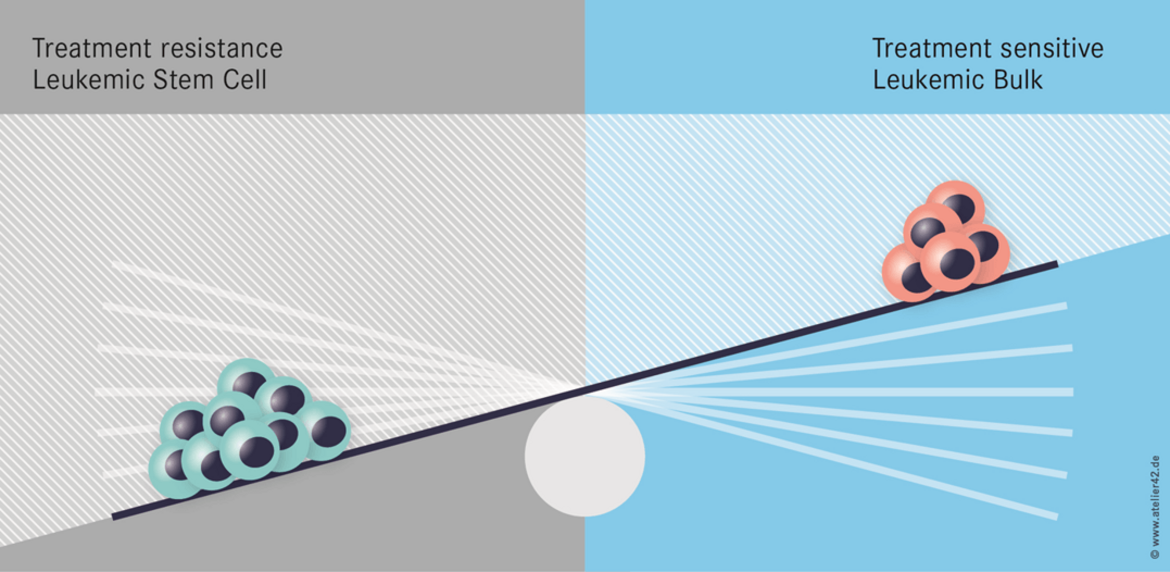
We have identified a novel epigenetic therapy resistance mechanism in Acute Myeloid Leukemia (Figure 2). Importantly, therapy resistance can be tackled by proteasome inhibition (or HSP90 inhibition). A multicenter clinical trial is planned to evaluate this concept in relapsed and refractory AML patients.
Chromatin and histone modifications as therapeutic targets are a field of active investigation. In an international collaborative effort we took part in the identification of LSD1 as a suitable target for differentiation therapy in AML (Schenk et al., 2012). Currently, we are performing a phase I/II clinical trial to test this approach in patients with refractory AML (NCT02261779). Biomarker analyses to identify why patients respond or do not respond are underway.
Alterations of DNA methylation occur frequently in most cancers. We are studying how changes in DNA methylation affect disease pathogenesis and therapy resistance in several cancer types. In genetic mouse models we are modeling leukemogenesis under conditions with increased or decreased DNA methyltransferase activity (Schulze et al., Blood 2016). In clinical trials we evaluate how DNA hypomethylating drugs such as Azacytidine can affect therapy response (Müller-Tidow et al., Leukemia 2016).
Mechanisms of stemness and aggressive disease in leukemia and beyond. In a clinical trial we are evaluating whether a novel CXCR4 inhibitor, BL-8040, that targets leukemic stem cells can improve therapy outcome for patients in first clinical remission https://clinicaltrials.gov/show/NCT02502968 .
With clinical trial data we developed a patient specific score to predict older AML patient´s response to induction chemotherapy (Krug et al., 2010) https://www.iblood.de/aml/.
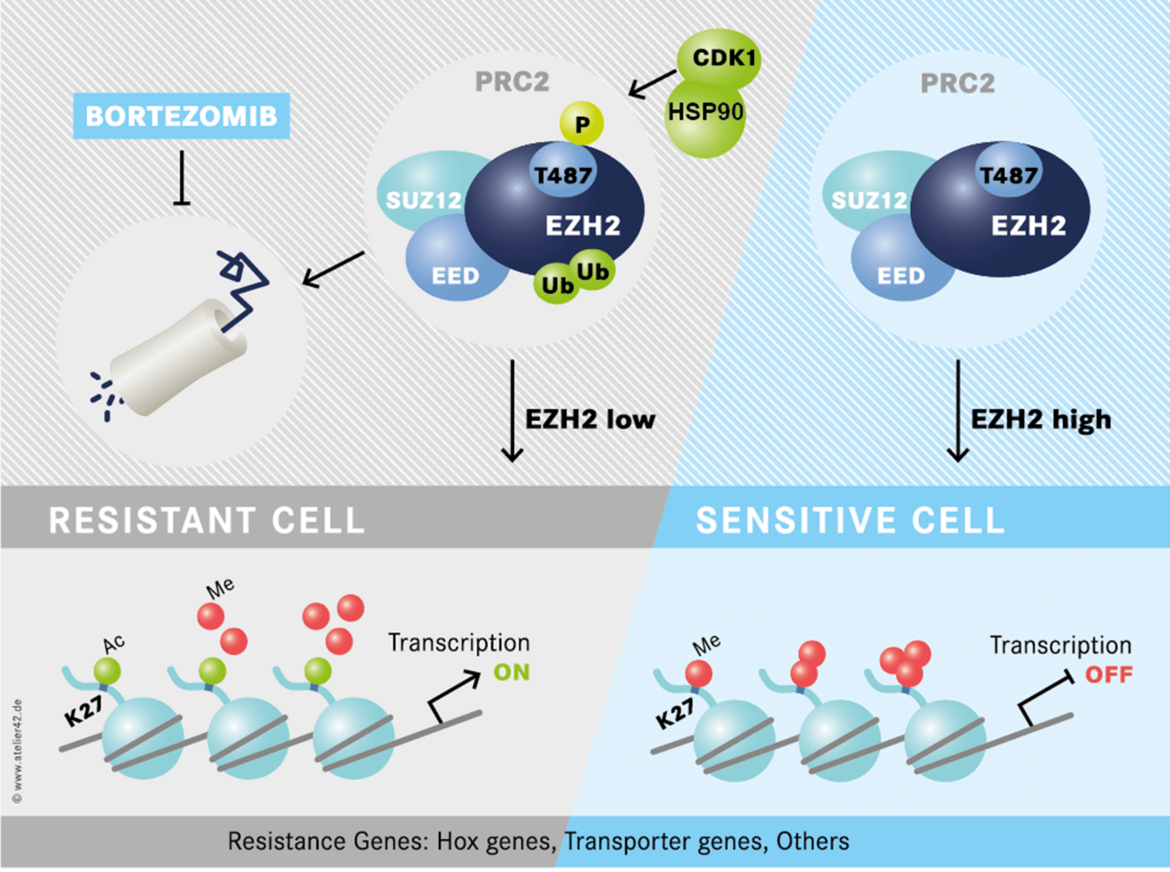
Recently, we identified a novel familial AML syndrome caused by germline DDX41 mutations (Figure 3) (Polprasat, Schulze et al., 2015).
These mutations predispose to AML, MDS and other hematological malignancies. We are currently investigating the pathogenetic mechanisms of mutations in the RNA helicase DDX41.
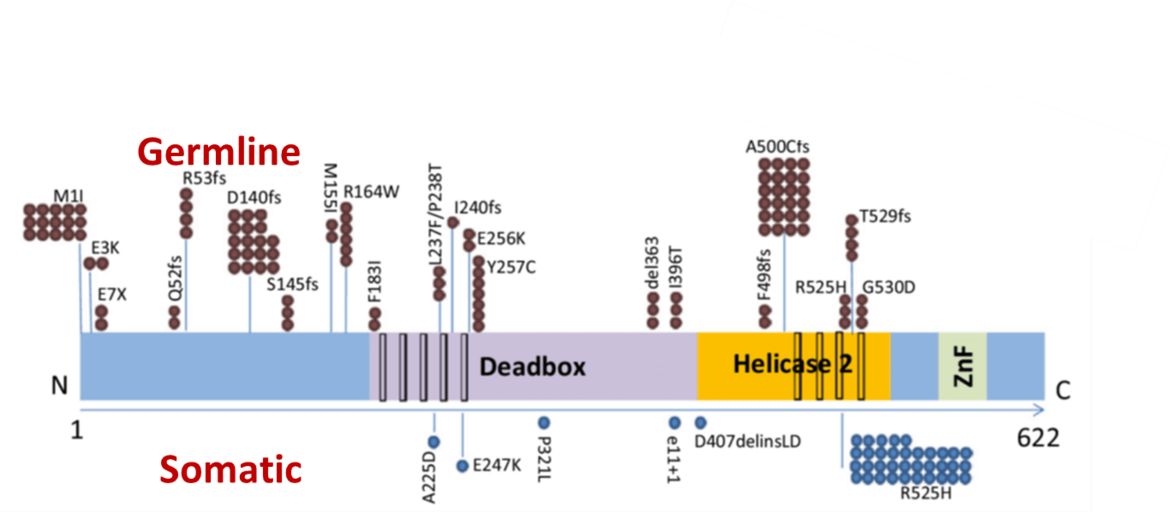
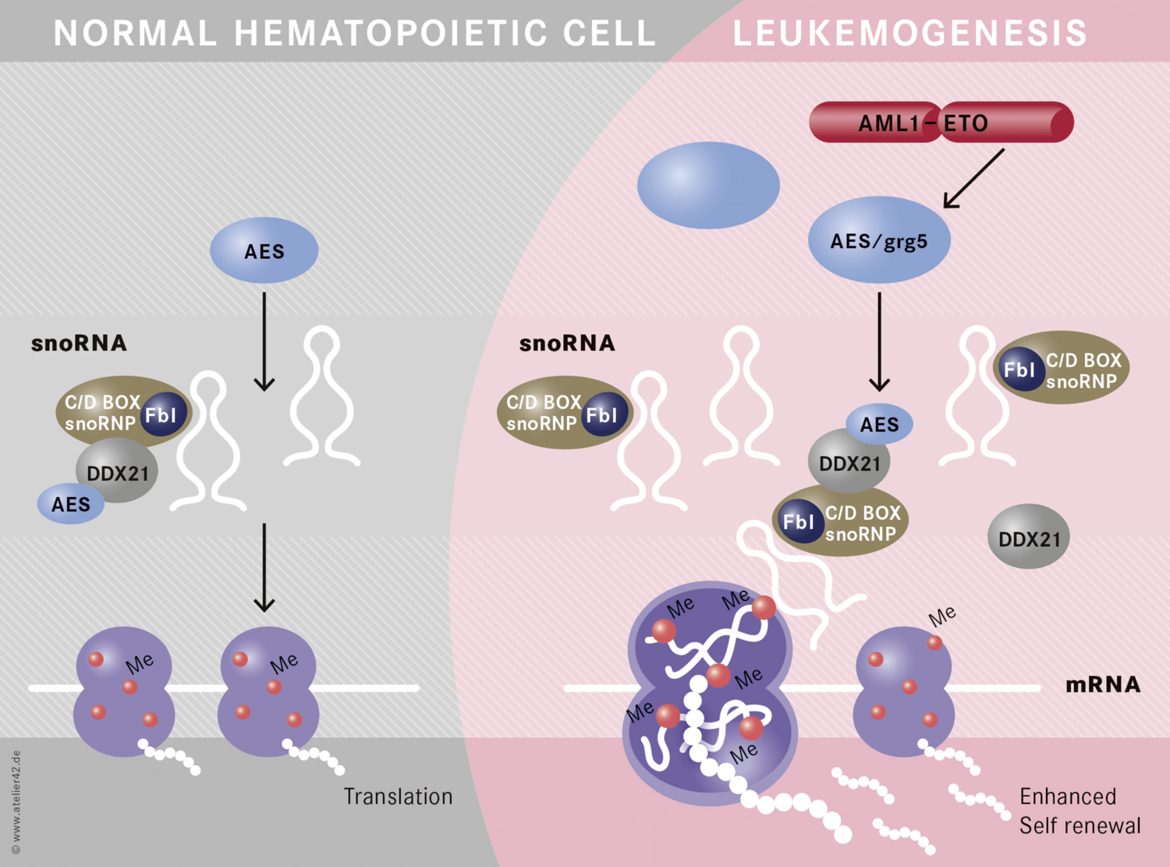
Non-coding RNAs are important mediators of disease aggressiveness. We discovered MALAT-1 (Metastasis-Associated Lung Adenocarcinoma Transcript 1) as a long non-coding RNA that is closely associated with lung cancer metastasis (Ji, Diederichs et al., 2003). Recently, we discovered that snoRNA functions are essential for leukemic stem cell self renewal in AML1-ETO induced leukemia (Figure 4)(Zhou et al., 2017). Currently, we are analyzing the specific functions of non-coding RNAs in leukemia and lung cancer.
Vacancies
We are always looking for outstanding postdocs, physician scientists and graduate students. If you are interested in our team, projects and research, please contact Carsten Müller-Tidow directly. Thank you very much!

NEWS
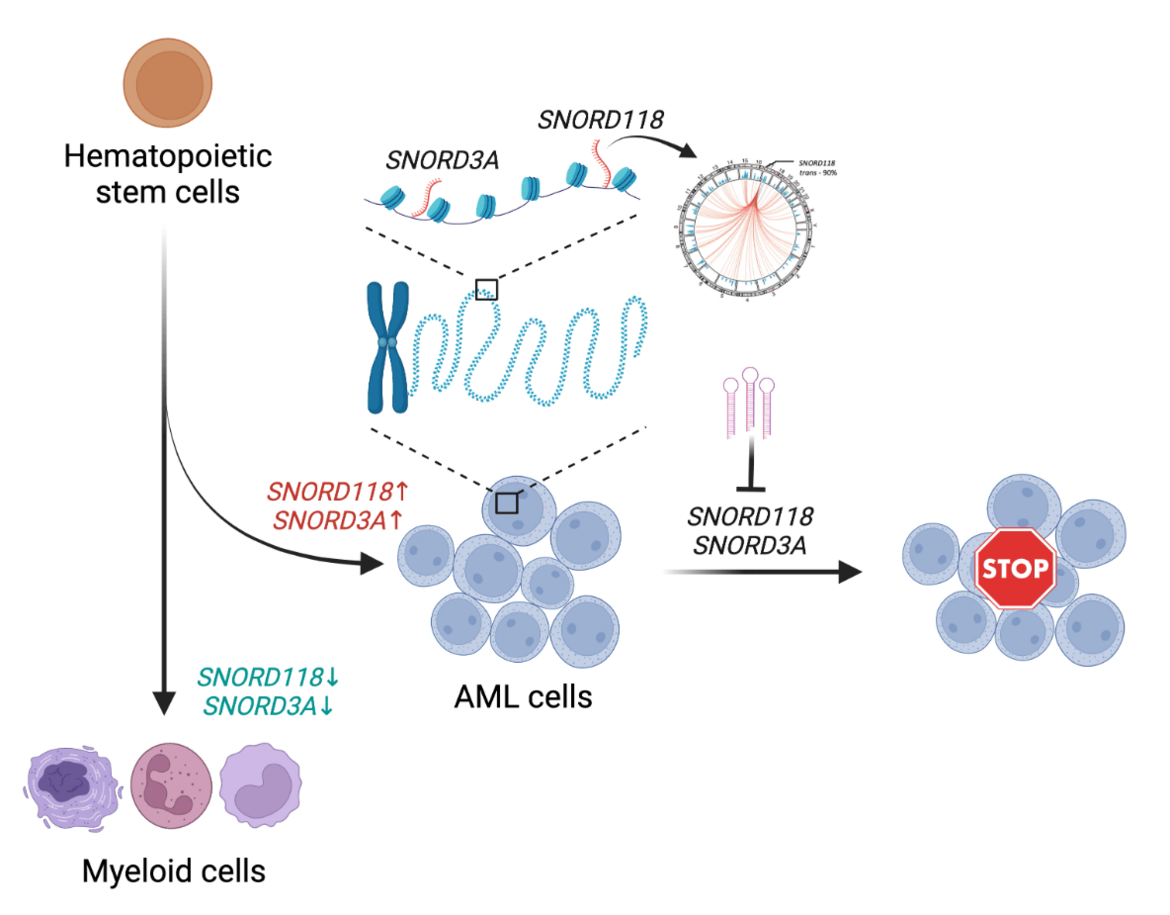
Unveiling the Hidden Regulators: Small Non-Coding RNAs in Leukemia
Published in Cancer, Cell & Molecular Biology, and Genetics & Genomics Jul 01, 2024
In the complex landscape of genetic regulation, the role of RNA goes far beyond its classical function as a mere messenger between DNA and protein synthesis. Recent research has illuminated the multifaceted roles of non-coding RNAs (ncRNAs) in transcription regulation, chromatin modulation and genome organization during tissue development as well as disease pathology [1,2]. Our study delves into this intricate world, uncovering the pivotal roles of chromatin-associated small non-coding RNAs in the maintenance and propagation of acute myeloid leukemia (AML). Continue reading.
Selected Publications
- Scheller, M., Ludwig, A.K., Göllner, S., Rohde, C., Krämer, S., Stäble, S., Janssen, M., Müller, J.A., He, L., Bäumer, N., Arnold, C., Gerß, J., Schönung, M., Thiede, C., Niederwieser, C., Niederwieser, D., Serve, H., Berdel, W.E., Thiem, U., Hemmerling, I., Leuschner, F., Plass, C., Schlesner, M., Zaugg, J., Milsom, M., Trumpp, A., Pabst, C., Lipka, D. B*. and Müller-Tidow, C.* (2021) Hotspot DNMT3A mutations in Clonal Hematopoiesis and Acute Myeloid Leukemia sensitize cells to Azacytidine via viral mimicry response. Nature Cancer, 2, 527-544
- Zhou F, Liu Y, Rohde C, Pauli C, Gerloff D, Köhn M, Misiak D, Bäumer N, Cui C, Göllner S, Oellerich T, Serve H, Garcia-Cuellar M-P, Slany R, Maciejewski JP, Przychodzen B, Seliger B, Klein H-U, Bartenhagen C, Berdel WE, Dugas M, Taketo MM, Farouq D, Schwartz S, Regev A, Hébert J, Sauvageau G, Pabst C, Hüttelmaier S, Müller-Tidow C: AML1-ETO requires enhanced snoRNP formation to induce self-renewal and leukemia. Nature Cell Biology 2017;19(7):844-855
- Göllner S, Oellerich T, Agrawal-Singh S, Schenk T, Klein HU, Rohde C, Pabst C, Sauer T, Lerdrup M, Tavor S, Stolzel F, Herold S, Ehninger G, Kohler G, Pan KT, Urlaub H, Serve H, Dugas M, Spiekermann K, Vick B, Jeremias I, Berdel WE, Hansen K, Zelent A, Wickenhauser C, Müller LP, Thiede C, Müller-Tidow C: Loss of the histone methyltransferase EZH2 induces resistance to multiple drugs in acute myeloid leukemia. Nature Medicine 2017;23(1):69-78.
- Schulze I, Rohde C, Scheller-Wendorff M, Bäumer N, Krause A, Herbst F, Riemke P, Hebestreit K, Tschanter P, Lin Q, Linhart H, Godley LA, Glimm H, Dugas M, Wagner W, Berdel WE, Rosenbauer F, Müller-Tidow C: Increased DNA methylation of Dnmt3b targets impairs leukemogenesis. Blood. 2016;127(12):1575-86.
- Müller-Tidow C*, Tschanter P*, Rollig C, Thiede C, Koschmieder A, Stelljes M, Koschmieder S, Dugas M, Gerss J, Butterfass-Bahloul T, Wagner R, Eveslage M, Thiem U, Krause SW, Kaiser U, Kunzmann V, Steffen B, Noppeney R, Herr W, Baldus CD, Schmitz N, Gotze K, Reichle A, Kaufmann M, Neubauer A, Schafer-Eckart K, Hanel M, Peceny R, Frickhofen N, Kiehl M, Giagounidis A, Gorner M, Repp R, Link H, Kiani A, Naumann R, Brummendorf TH, Serve H, Ehninger G, Berdel WE, Krug U, Study Alliance Leukemia G: Azacitidine in combination with intensive induction chemotherapy in older patients with acute myeloid leukemia: The AML-AZA trial of the Study Alliance Leukemia. Leukemia. 2016;30(3):555-61.
- Polprasert C*, Schulze I*, Sekeres MA, Makishima H, Przychodzen B, Hosono N, Singh J, Padgett RA, Gu X, Phillips JG, Clemente M, Parker Y, Lindner D, Dienes B, Jankowsky E, Saunthararajah Y, Du Y, Oakley K, Nguyen N, Mukherjee S, Pabst C, Godley LA, Churpek JE, Pollyea DA, Krug U, Berdel WE, Klein HU, Dugas M, Shiraishi Y, Chiba K, Tanaka H, Miyano S, Yoshida K, Ogawa S, Müller-Tidow C*, Maciejewski JP*: Inherited and Somatic Defects in DDX41 in Myeloid Neoplasms. Cancer Cell. 2015;27(5):658-70.
- Bäumer S, Bäumer N, Appel N, Terheyden L, Fremerey J, Schelhaas S, Wardelmann E, Buchholz F, Berdel WE, Müller-Tidow C: Antibody-mediated delivery of anti-KRAS-siRNA in vivo overcomes therapy resistance in colon cancer. Clin Cancer Res. 2015;21(6):1383-94.
- Hascher A*, Haase AK*, Hebestreit K, Rohde C, Klein HU, Rius M, Jungen D, Witten A, Stoll M, Schulze I, Ogawa S, Wiewrodt R, Tickenbrock L, Berdel WE, Dugas M, Thoennissen NH*, Müller-Tidow C*: DNA methyltransferase inhibition reverses epigenetically embedded phenotypes in lung cancer preferentially affecting polycomb target genes. Clin Cancer Res. 2014;20(4):814-26.
- Schoofs T*, Rohde C*, Hebestreit K, Klein HU, Göllner S, Schulze I, Lerdrup M, Dietrich N, Agrawal-Singh S, Witten A, Stoll M, Lengfelder E, Hofmann WK, Schlenke P, Buchner T, Hansen K, Berdel WE, Rosenbauer F, Dugas M, Müller-Tidow C: DNA methylation changes are a late event in acute promyelocytic leukemia and coincide with loss of transcription factor binding. Blood. 2013;121(1):178-87.
- Schenk T, Chen WC, Göllner S, Howell L, Jin L, Hebestreit K, Klein HU, Popescu AC, Burnett A, Mills K, Casero RA, Jr., Marton L, Woster P, Minden MD, Dugas M, Wang JC, Dick JE, Müller-Tidow C, Petrie K, Zelent A: Inhibition of the LSD1 (KDM1A) demethylase reactivates the all-trans-retinoic acid differentiation pathway in acute myeloid leukemia. Nat Med. 2012;18(4):605-11.
- Krug U, Rollig C, Koschmieder A, Heinecke A, Sauerland MC, Schaich M, Thiede C, Kramer M, Braess J, Spiekermann K, Haferlach T, Haferlach C, Koschmieder S, Rohde C, Serve H, Wormann B, Hiddemann W, Ehninger G, Berdel WE, Büchner T*, Müller-Tidow C*, German Acute Myeloid Leukaemia Cooperative G, Study Alliance Leukemia I: Complete remission and early death after intensive chemotherapy in patients aged 60 years or older with acute myeloid leukaemia: a web-based application for prediction of outcomes. Lancet 2010;376(9757):2000-8.
- Ji P*, Diederichs S*, Wang W, Boing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, Müller-Tidow C. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003;22(39):8031-8041.